Result & SNP List
Cladefinder nach YSEQ
Range I-YSC261 Ytree
Visigothic Hypothesis
Tree after Morley
Haplogroups A0-T, A1, BT, CF, CT
Haplogroup E (M96)
🧬 Was bedeutet es, wenn Marker anderer Haplogruppen auftauchen?
Haplogroup F (M89)
Haplogroup I1 (M253)
Neolithic "Nuragic" Markers: Haplogroup I2 (L672/S327, P222/U250/S118)
Markers from J2a (L152)
Markers from E1: Subclads E1b1a1a1 (L88/L88.1), E1b1a1a1h (P-269)
1900-950 BCE: I1 related tree for latest marker S1954/YSC0000261 (Z140/S440, S1953/Z2535 → I1a2a1a1a1a)
1300 BCE - 700 CE: E/R related tree hypotheses for latest marker S1954/YSC0000261
Examining: I1a2a1a1a~1 [I1-L41] [I1a2a1a1a1, I1-S1953/Z2535, with admixtures from I2, J2a, E] (ytree.morleydna.com/extractFromAutosomal)
"Some autosomal genetic genealogy tests (such as 23andMe, AncestryDNA and MyHeritage - but not Family Finder) also contain a few hundred Y-DNA markers. The Y-DNA data from these tests is of lower quality, but may still suffice for a very, very general Y-DNA haplogroup classification", Creative Commons Licence CC BY-NC-ND 3.0, ytree.morleydna.com.
"This suggested classification does not account for the following positive SNPs:
Most specific position on the YFull YTree is I-YSC261
I-YSC261
BY1910/ZS11584(?) DF77/S1969(?) Y125368($) Y4016(?) Y488283(?) Y489121(?) Y489123(?) Y489124(?) Y489210(?) YSC261/S1954/YSC0000261+ Z4727/S1971($) Z4728/S1960(?) Z4729/S1987(?) Z4730/S1968(?) Z4731/S1962(?) Z4732($) Z4733($) Z4734(?) Z4735(?) Z4736/S1978($) Z4737(?)
┗━ I-L338
L338/S197
"Available Panels: YSEQ recommends the I1-Z140 Panel Predicted I-YSC261 is downstream of the panel root. This panel may be applicable if it tests subclades below I-YSC261. Please verify and check with YSEQ customer support."
Cl 95% 4.100 - 2.700 ybp.
Next best prediction (scored 103 compared to 104) I-Z2535.
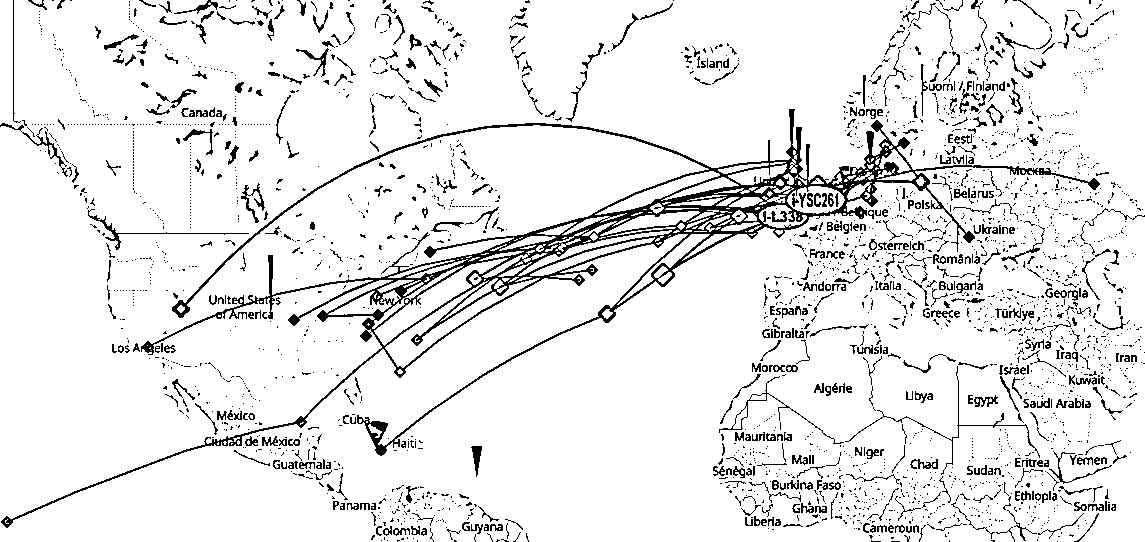 Range I-YSC261 Ytree (unter Auslassung von USA; wo nicht anders vermerkt, handelt es sich um moderne Samples):
Range I-YSC261 Ytree (unter Auslassung von USA; wo nicht anders vermerkt, handelt es sich um moderne Samples):
1. I-YSC261 (4.000 ybp): Syddanmark / Dänemark (DK = Ancient sample Id: CGG106777, age: 1050; 900 - 1.200 ybp)
1.1. I-L338 (3.400 ybp): Schottland, Galloway/Schottland (I-A1968, 850 ybp); Saint Thomas / Jamaika (I-FT288387, 850 ybp); Středočeský kraj / Czechia, Noord-Holland (I-A2398, 2.900 ybp); UK (I-BY31701, 1.600 ybp)
1.2. I-A12974 (2.900 ybp): Gateshead / UK (I-A12974*, 2.900 ybp); Herfordshire / UK (I-A12720, 275 ybp); Akershus / Norwegen (I-8673*, 1.350 ybp; I-8674, 500 ybp); Dnipropetroovs'ka Oblast' / Ukraine, Botoşani / Rumänien, Mazowieckle / Polen (I-Y12663, 1.350 ybp)
1.3.1. I-S12289: Nieder�sterreich (I-S12289*, 2.900 ybp; Ancient Sample Id: MLE004, age: 1.200; 1.150 - 1.250 ybp); Västernorrlands / Schweden (I-A20048, 2.800 ybp); Staffordshire / UK (I-A11323, 900 ybp); Wittshire / UK (I-FGC60623, 2.500 ybp); Derbyshire / UK (I-Y252117, 2.500 ybp; I-Y3636, 1.300 ybp; I-A4638, 1.250 ybp)
1.3.2. I-A4586 (2.800 ybp): Nordrhein-Westfalen (I-BY51406, 2.600 ybp); Cambridgeshire / UK (I-A21901*, 1.800 ybp); Skåne län / Schweden (I-Y197065, 1.300 ybp); Västra Götalands län / Schweden (I-FT354407, 1.300 ybp)
1.3.3. I-A20033 (2.800 ybp): Lanarkshire / Schottland (I-A20451*, I-A20831, 750 ybp); Lanarkshire, Dunbartonshire, Schottland (I-Y131422, 750 ybp); Zuid-Holland (I-A20032, 750 ybp; I-BY32573, I-A20047*, I-A20044, I-A22686*, 500 ybp; I-FGC29004, 400 ybp); England, Zuid-Holland (A22865 = S22865, 400 ybp)
1.3.4. I-Y14670 (2.800 ybp): Mecklenburg, Akershus / Norwegen, Haut-Rhin / Frankreich, Syddanmark / Dänemark (I-Y8334, 2.300 ybp; DK = Ancient Sample Id: VK327, age: 1.000; 850 - 1.150); Skåne län / Schweden, Ontario / Kanada (I-Y322267*, 1.750 ybp); Ul'yanovskaya Oblast / Russland (I-PH1410, 1.750 ybp); Wales (I-Y185625*, 1.750 ybp); New South Wales / Australia, Massachusetts (I-A7320, 1.200 ybp); Barbados, Virginia (I-A4637, 1.750 ybp); Utrecht (I-PH4462, 1.750 ybp); Dorset / GB, Irland (I-A18483, 1.550 ybp); D�nemark (I-BY50971, 1.550 ybp)
1.3.5. I-Y4015 (2.800 ybp): Dalamas län / Schweden, England, Baden-W�rttemberg, Hovedstaden / Dänemark (DK = 1.749; 1550 -1949 ybp); UK (I-A1845, 2.500 ybp); Östergötlands (I-A11569, 2.500 ybp); Nordylland / DK (I-Y132919*, 2.500 ybp); Niedersachsen (I-Y132926, 2.500 ybp); Bremen / DE (I-A16200, 2.00 ybp); England (I-FT111200, 800 ybp); England, Saint Michael / Barbados (I-FT111200, 800 ybp); Limburg / NL (I-A8595*, 2.500 ybp); Gloucestershire, England (I-A8596*, 2.400 ybp; I-FT 409631, 325 ybp); England (I-A1627*, 2.300 ybp); Invernessshire / Schottland (I-A1626*, 1.700 ybp); Barbados (I-BY31695*, 2.300 ybp); England (I-A1790, 1.750 ybp); UK (I-S1972, 1.750 ypb); UK (I-A9538, 1.050 ybp); Schottland (I-YFS523700, 700 ybp); Irland (I-Y23905*, 700 ybp); Antrim / UK (I-Y219852, 350 ybp); Lanarkshire / Schottland (I-FGC74328*, 350 ybp); Irland (I-FGC74329, 350 ybp); Schottland (I-FS273194, 350 ybs); Dublin / Irland (I-A264, 700 ybp); Irland (I-YFS2353752).
[ yfull.com ], [ hras.yseq.net ], [ phylogeographer.com ]
The Visigothic-Iberian Hypothesis and the I1 migration from Scandinavia to Poland as Wielbark culture before Gothic ethnogenesis
"The Wielbark culture (German: Wielbark-Willenberg-Kultur; Polish: Kultura wielbarska) is an Iron Age archaeological complex that flourished on the territory of today's Poland from the 1st century to the 5th century. The Wielbark culture is associated with the Goths and related Germanic peoples, and played an important role in the Amber Road." (WP).
(1) Ireneusz Stolarek et al.: "Genetic history of East-Central Europe in the first millennium CE", in: "Genome Biology" 24, Article 173, 2023 [ link.springer.com ]:
"The detected Y-hgs (155 in total [159 samples]) belonged to the following major branches of the Y-chromosome phylogenetic tree: E, F, G2a, I1, I2a, J2a, J2, N1a, R1a, and R1b (Fig. 3a, Additional file 2: Table S3). The Y-hg frequencies in the IA [Iron Age] and MA [Middle Ages] groups were significantly different (Fisher's exact test, p < 0.001). In the IA group, the Y-hg I1/I1a was the most frequent [13x I1, 6x I1a, 6x R1b, 5x G2a, 4x R1a, 11x Other, 1x Unk; Locations: Czarnowko, Kowalewko, Maslomecz, Pruszcz Gdanski], whereas in the MA group [66x R1a, 7x R1b, 6x I2a, 5x R1, 5x E1b, 2x I1, 2x I1a, 17x Other, 3x Unk; Locations: Balczewo, Dziekanowice, Golun, Konskie, Lad, Legowo, Markowice, Milicz, Niemcza, Oblarczkowo, Ostrow Lednicki, Plonsk, Poznan, Rumin, Santok], R1a predominated (Fig. 3c). The frequencies of both haplogroups underwent major changes between the two periods: the I1/I1a frequency decreased from 41.3% in the IA to 3.5% in the MA, whereas the R1a frequency increased from 8.6% in the IA to 57.5% in the MA. In the IA I1 individuals, the highest frequency was observed for the L118 mutation (41.3%), which is common in present-day Scandinavia and Finland. [...] The genomes of IA individuals with the Y-hg I1 were similar to those observed in contemporary Northwestern Europeans, whereas the genomes of MA males with the Y-hg I1 were dominated by contemporary East-Central European components".
(2) Olade et al.: "The genomic history of the Iberian Peninsula over the past 8000 years", in: "Science", 15 Mar 2019, Vol 363, Issue 6432, pp. 1230-1234 [ pmc.ncbi.nlm.nih.gov ]:
"We assembled genome-wide data from 271 ancient Iberians of whom 176 are from the largely unsampled period after 2000 BCE, thereby providing a high resolution time transect of the Peninsula. We document high genetic substructure between northwestern and southeastern hunter-gatherers prior to the spread of farming. We reveal sporadic contacts between Iberia and North Africa by ~2500 BCE, and by ~2000 BCE the replacement of 40% of Iberia's ancestry and nearly 100% of its Y-chromosomes by people with Steppe ancestry. In the Iron Age, we show that Steppe ancestry had spread not only into Indo-European-speaking regions but also into non-Indo-European-speaking ones, and we reveal that present-day Basques are best described as a typical Iron Age population without the admixture events that later impacted the rest of Iberia. Beginning at least in the Roman period, we document how the ancestry of the Peninsula was transformed by gene flow from North Africa and the eastern Mediterranean".
Table S4: 78x R1, 53x I2, 19x G2, 14x I = 3x NE_Iberia_MLN + 2x SE_Iberia_MLN + SW_Iberia_CA + NE_Iberia_c.6CE_PL + SE_Iberia_c.5-8CE + 3x C_Iberia_CA + 2x SE_Iberia_CA + SW_Iberia_CA, 13x E1 = SE_Iberia_c.10-16CE_Afr + 3x SE_Iberia_c.5-8CE + C_Iberia_CA_Afr + NE_Iberia_c.8-12CE + NE_Iberia_c.6CE_PL + 6x SE_Iberia_c.10-16CE, 11x CT, 6x J = 2x NE_Iberia_Hel (Empúries2) + NE_Iberia_RomP (Empúries2) + SE_Iberia_c.3-4CE + SE_Iberia_c.5-8CE + SE_Iberia_c.10-16CE, 4x J2 = NE_Iberia_c.6CE_PL + 2x SE_Iberia_c.10-16CE + SE_Iberia_c.3-4CE, 4x F, 3x R = 2x NE_Iberia_Greek (Empúries1) + NE_Iberia_RomP, 20x Other; n = 226, CA = Copper Age, PL = Post-Levant, Hel = Hellenistic Period, RomP = Roman Period, MLN = Middle Late Neolithic, 3.500 - 2.500 v.chr.Z.
(3) Adams et al.: "The Genetic Legacy of Religious Diversity and Intolerance: Paternal Lineages of Christians, Jews, and Muslims in the Iberian Peninsula", in: "American Journal of Human Genetics", 2008 Dec 5, Vol. 83, Issue 6, pp. 725-736, [ doi: 10.1016/j.ajhg.2008.11.007 ].
Figure 1 shows for "All Iberian Peninsula", n = 1.140 (Aragon, n = 34; Andalusia East, n = 95; Andalusia West, n = 73; Asturias, n = 20; Basque Country, n = 116; Castilla la Mancha, n = 63; Castile NE, n = 31; Castile NW, n = 100; Catalonia, n = 80; Extremadura, n = 52; Galicia, n = 88; Gascony, n = 24; Portugal North, n = 60; Portugal South, n = 78; Valencia, n = 73; Majorca, n = 62; Minorca, n = 37; Ibiza, n = 54), "[b]elow the phylogeny are given the percentages of chromosomes carrying the observed haplogroup", 2 (0.2%) E1, 3 (0.3%) E3a, 11 (1%) E3b*, 42 (4%) E3b1, 49 (4%) E3b2, 11 (1%) E2b3, 1 (0.1%) F*(xG-K), 57 (5%) G, 68 (6%) I, 14 (1%) J(xJ2), 88 (8%) J2, 29 (3%) K*, 2 (0.2%) Q*, 1 (0.1%) R1*, 14 (1%) R1a1, 2 (0.2%) R1b*, 623 (55%) R1b3*, 2 (0.2%) R1b3b, 37 (3%) R1b3d, 84 (7%) R1b3f, - R2.
For "Aragon ARA", n = 34, 3% E3b*, 3% E3b2, 18% I, 12% J2, 6% K*, 3% R1a1, 38% R1b3*, 3% R1b3d, 15% R1b3f.
For "Catalonia CAT", n = 80, 1% E3b1, 1% E3b2, 1% E2b3, 6% G, 3% I, 6% J2, 59% R1b3*, 1% R1b3d, 21% R1b3f.
For "Basque Country, BAS", n = 116, 1% E3b2, 8% I, 1% J(xJ2), 3% J2, 1% Q*, 63% R1b3*, 2% R1b3b, 13% R1b3d, 9% R1b3f.
For "Portugal North NPO", n = 60, 3% E1, 2% E3b*, 5% E3b1, 3% E3b2, 2% E2b3, 12% G, 2% I, 2% J(xJ2), 7% J2, 2% K*, 3% R1a1, 47% R1b3*, 2% R1b3d, 10% R1b3f.
For "Portugal South SPO", n = 78, 1% E3a, 3% E3b*, 4% E3b1, 8% E3b2, 1% E2b3, 9% G, 4% I, 3% J(xJ2), 15% J2, 5% K*, 1% R1a1, 44% R1b3*, 3% R1b3f.
For "Ibiza, IBZ", n = 54, 4% E3b1, 4% E2b3, 13% G, 2% I, 4% J2, 17% K*, 46% R1b3*, 4% R1b3d, 7% R1b3f.
For "All N.Africa", n = 361 (Morocco, n = 147; Algeria, n = 46; Tunisia, n = 139; Saharawi, n = 29), 5 (1%) E1, 13 (4%) E3a, 13 (4%) E3b*, 21 (6%) E3b1, 196 (54%) E3b2, 5 (1%) E2b3, 11 (3%) F*(xG-K), - G, 1 (0.3%) I [Morocco], 63 (17%) J(xJ2), 10 (3%) J2, 3 (1%) K*, - Q*, - R1*, - R1a1, - R1b*, 18 (5%) R1b3*, - R1b3b, - R1b3d, 1 (0.3%) R1b3f, - R2.
For "Sephardic Jews", n = 174, - E1, 1 (1%) E3a, 2 (1%) E3b*, 6 (3%) E3b1, 1 (1%) E3b2, 6 (3%) E2b3, - F*(xG-K), 27 (16%) G, 1 (1%) I, 38 (22%) J(xJ2), 43 (25%) J2, 11 (6%) K*, 4 (2%) Q*, - R1*, 8 (5%) R1a1, 2 (1%) R1b*, 20 (11%) R1b3*, 4 (2%) R2.
Figure 4, and Table S2, show "Iberian, North African, and Sephardic Jewish Admixture Proportions among Iberian Peninsula Samples":
Population "Aragon" with 59.4% mY1 Basque Ancestry Proportion at 11.6% SD, with 4.8% Moroccan Ancestry Proportion at 3.6% SD, with 35.8% Sephardic Ancestry Proportion at 12.0% SD.
Population "Asturias" with 44.3% mY1 Basque Ancestry Proportion at 15.3% SD, with 10.5% Moroccan Ancestry Proportion at 7.3% SD, with 45.2% Sephardic Ancestry Proportion at 16.2% SD [n = 20!].
Population "Catalonia" with 91.5% Basque Ancestry Proportion at 7.0% SD, with 2.3% Moroccan Ancestry Proportion at 2.5% SD, with 6.2% Sephardic Ancestry Proportion at 7.1% SD.
Population "Castile, NW" with 65.4% Basque Ancestry Proportion at 6.0% SD, with 21.7% Moroccan Ancestry Proportion at 5.4% SD, with 12.9% Sephardic Ancestry Proportion at 2.7% SD.
Population "Extremadura" with 52.3% Basque Ancestry Proportion at 9.4% SD, with 19.0% Moroccan Ancestry Proportion at 7.1% SD, with 28.7% Sephardic Ancestry Proportion at 10.4% SD.
Population "Galicia" with 62.3% Basque Ancestry Proportion at 7.4% SD, with 20.8% Moroccan Ancestry Proportion at 5.6% SD, with 16.9% Sephardic Ancestry Proportion at 7.5% SD.
Population "Portugal North" with 64.7% mY1 Basque Ancestry Proportion at 8.8% SD, with 11.8% Moroccan Ancestry Proportion at 5.3% SD, with 23.6% Sephardic Ancestry Proportion at 9.1% SD.
Population "Portugal South" with 47.6% mY1 Basque Ancestry Proportion at 8.1% SD, with 16.1% Moroccan Ancestry Proportion at 6.1% SD, with 36.3% Sephardic Ancestry Proportion at 9.2% SD.
Population "Ibiza" with 63.2% mY1 Basque Ancestry Proportion at 9.0% SD, with 3.8% Moroccan Ancestry Proportion at 4.3% SD, with 33.0% Sephardic Ancestry Proportion at 9.7% SD.
(4) Conversion Studies
a. Norman Golb: "Jewish Proselytism. A Phenomenon in the Religious History of Early Medieval Europe", Cincinatti: Judaic Studies Program 1987 [uchicago.edu ].
b. Sabrina Sp�th: "Konversionen auf der mittelalterlichen Iberischen Halbinsel. Eine vergleichende Betrachtung dreier Konvertiten im Spiegel der Quellen", Erlangen: FAU University Press 2016 [ library.oapen.org ].
(5) Elizabeth Caldwell Hirschman and Donald Panther-Yates: "Toward a Sephardic Haplogroup Profile in the New World", Longmont, Colorada: DnaConsultants.Com 2006.
Table 29, "Summary of Sephardic Y-Haplotype Distribution":
| Canary Islands | Azores | Cuba | Puerto Rico | Mexico | New Mexico | |
|---|---|---|---|---|---|---|
| n | 34 | 13 | 44 | 67 | 129 | 142 (62) |
| Haplogroup | ||||||
| R1b | 55.9 | 61.5 | 72.7 | 49.3 | 55.8 | 55.6 (56.1) |
| E3b | 17.6 | 0.0 | 9.1 | 12.0 | 11.6 | 9.5 (4.5) |
| I, I1c, I1b | 8.8 | 30.8 | 9.1 | 13.4 | 12.4 | 7.9 (15.2) |
| J, J1, J2 | 0.0 | 0.0 | 2.3 | 12.0 | 11.5 | 18.3 (18.2) |
| G, G2 | 8.8 | 7.7 | 2.3 | 4.5 | 5.4 | 3.2 (4.5) |
| K2 | 2.9 | 0.0 | 0.0 | 6.0 | 1.5 | 0.0 (0.0) |
| O3 | 5.9 | 0.0 | 0.0 | 0.0 | 0.7 | 0.0 (0.0) |
| R1a1 | 0.0 | 0.0 | 0.0 | 3.0 | 0.0 | 0.7 (1.5) |
 Surname lists excluded loc. cit. (New Mexico has the surname "Aragon"). Sephardic Jews, primarily fleeing the Portuguese Inquisition in Recife, Brazil, established a significant community in Barbados around 1654 (Speightstown Synagogue). In 1750 "Sephardic Jews account for 800 out of a total (white) population of 12,000. Over 400 sugar-processing windmills dot the landscape of Barbados" [ jewishbarbados.org ].
Surname lists excluded loc. cit. (New Mexico has the surname "Aragon"). Sephardic Jews, primarily fleeing the Portuguese Inquisition in Recife, Brazil, established a significant community in Barbados around 1654 (Speightstown Synagogue). In 1750 "Sephardic Jews account for 800 out of a total (white) population of 12,000. Over 400 sugar-processing windmills dot the landscape of Barbados" [ jewishbarbados.org ].
Lar Gilinsky: "Jews by Haplotype, Likeness and History. A nuanced illustration of relatedness", humgeno.org, 2023: "Sephardic marriage records of Amsterdam: The following are 79 references regarding the 'New World' in previous centuries: Curacao 47, Suriname 19, Barbados 10, Kingston 1, Port St. Marten 1, Saint Croix 1" (S. 62).
"Sephardic Jews have a higher proportion of haplogroups R1b (29.5%, compared to 11.4% among the Ashkenazic Jews) and I (11.5%, compared to 4% among the Ashkenazic Jews). This is unsurprising as these two haplogroups are found in the highest frequency in Atlantic Europe and the Balkans, respectively, two areas where Sephardic but not Ashkenazic Jews settled in large numbers. Ashkenazic Jews, in contrast, have a higher frequency of haplogroups J (43% compared to 28.2% among Sephardic Jews) and E1b1b (22.8% compared to 19.2% among Sephardic Jews), which have been carried on since pre-Diaspora times (Haplogroup J is most common in the Middle East and haplogroup E1b1b is widespread in the Horn of Africa.) Such patterns further support the abovementioned generalization that Ashkenazic Jews remained more isolated from their host populations than Sephardic Jews. (These data are from Nebel, Filon, Brinkmann, Majumder, Faerman, and Oppenheim, 'The Y Chromosome Pool of Jews as Part of the Genetic Landscape of the Middle East', American Journal of Human Genetics 2001, 69(5): 1095-1112.) The upshot of this discussion is that there is no haplogroup or mutation that unambiguously identifies Sephardic Jews" (esefarad.com/who-counts-as-a-sephardic-jew).
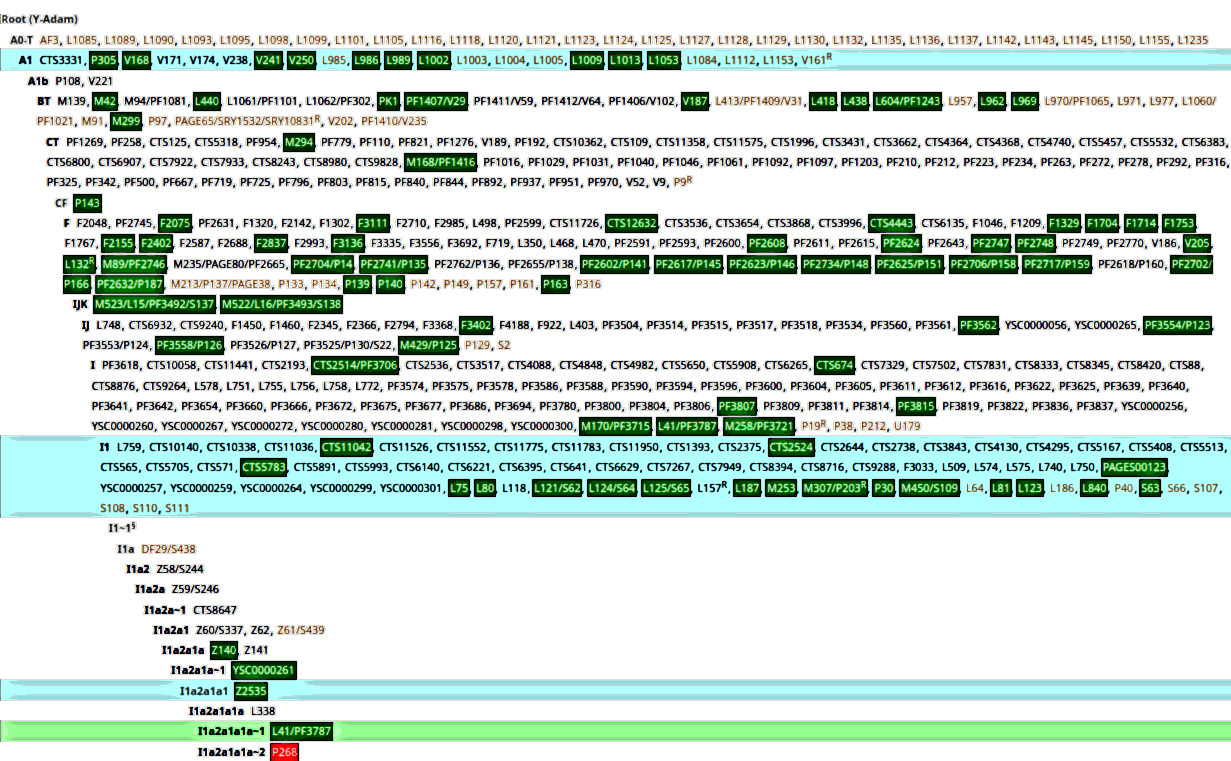
Haplogroups A0-T, A1, BT, CF, CT
WP: "Haplogroup A-L1085, also known as haplogroup A0-T is a human Y-DNA haplogroup. It is part of the paternal lineage of almost all humans alive today", "Possible time of origin 140,000 YBP, 125,000 - 382,000 YBP. Possible place of origin Central-Northwest Africa".
"Haplogroup A-P305 also known as A1 is a human Y-chromosome DNA haplogroup. Like its parent haplogroup haplogroup A0-T (A-L1085), A1 includes the vast majority of living human males. It emerged in Africa approximately 161,300 years ago. By comparison, members of its sole sibling subclade, haplogroup A0 - the only other primary subclade of haplogroup A0-T - are found mostly in Africa" (yfull.com/tree/A1), "Possible time of origin 161,300 years BP. Possible place of origin Africa" (WP).
"Haplogroup BT M91, also known as Haplogroup A1b2 (and formerly as A4, BR and BCDEF), is a Y-chromosome haplogroup. [...] Basal BT* has not been documented in any living individuals or ancient remains. No definite examples of BT(xCF,DE) - i.e. members of BT outside the only two known branches of CT, namely haplogroups CF and DE - have been identified", "Possible time of origin 150,000-145,000 BP. Possible place of origin Africa" (WP).
"Haplogroup CF, also known as CF-P143 and CT(xDE), is a human Y-chromosome DNA haplogroup. CF is defined by the SNP P143, and its existence and distribution are inferred from the fact that haplogroups descended from CF include most human male lineages in Eurasia, Oceania, and The Americas. CF descends from CT (CT-M168), and is the sibling of DE. CF has two basal branches, Haplogroup C and Haplogroup F", "Possible time of origin 75,000-70,000 BP. Possible place of origin: Africa" (WP).
"Haplogroup E-M96 is a human Y-chromosome DNA haplogroup. It is one of the two main branches of the older and ancestral haplogroup DE [descending from CT], the other main branch being haplogroup D. The E-M96 clade is divided into two main subclades: the more common E-P147, and the less common E-M75", "Possible time of origin 65,200 years BP, 69,000 years BP, or 73,000 years BP. Coalescence age 52,300 years BP. Possible place of origin East Africa, West Africa, or Eurasia" (WP).
- CT
- DE (M1/YAP, M145/P205, M203, P144, P153, P165, P167, P183)
- E (M96)
- E1 (P147)
- E1a (M33, M132, "formerly E1", WP)
- E1a1 (M44, "mostly found in Europe, West Asia and among Ashkenazi Jews", WP)
- E1a2 (Z958, "exclusively identified in Sub-Saharan Africa", WP)
- E1b (P177)
- E1b1 (P2, DYS391p, "formerly E3", WP)
- E1b1a (V38)
- E1b1a1 (M2, +20 Segm. see above, "[f]ound in Africa, especially among Niger-Congo-speaking populations; formerly E3a", WP)
- E1b1a2 (M329, "[f]ound in Africa, especially in Ethiopia among Omotic-speaking populations; formerly E3*", WP)
- E1b1b (M215, "formerly known as E3b", "Phönizier")
- E1b1b1 (M35, "[f]ound in Horn of Africa, North Africa, the Middle East, and Europe (esp. in areas near the Mediterrian and the Balkans); formerly E3b")
- E1b1b1a (V68)
- E1b1b1b (Z827)
- E1b1b1b1 (Z830, L19, V257)
- E-M81 ("Die Untergruppe E-M81 [E3b2] ist vorherrschend unter nordafrikanischen Berber sprechenden Bevölkerungen und Maghrebinern")
- E-M123 ("Possible time of origin 15,000-20,000 ybp. Possible place of origin Middle East")
- E-M34 ("In fact E-M34 seems to be restricted to Semitic speakers in Ethiopia, as it has not been detected in other populations in the region such as Somalia, Kenya (Cruciani et al. 2004)", WP)
- E1b1b1b2 (L19)
- E1b1b1b1 (Z830, L19, V257)
- L539 ("Around 23,900 years before present (YBP), the paternal lineage of E-L539 took shape as it diverged from the common ancestor E-M35 branch. The most recent common ancestor (TMRCA) of this branch is estimated to have been born around 19,000 years before present (YBP)", somaliy.com/Branch/E-L539)
- E-M78 (yfull.com/tree/e-m78) [E3b1, "Balkan"]
- E1b1b1 (M35, "[f]ound in Horn of Africa, North Africa, the Middle East, and Europe (esp. in areas near the Mediterrian and the Balkans); formerly E3b")
- E1b1a (V38)
- E1b1 (P2, DYS391p, "formerly E3", WP)
- E1a (M33, M132, "formerly E1", WP)
- E2 (M75)
- E1 (P147)
- E (M96)
- DE (M1/YAP, M145/P205, M203, P144, P153, P165, P167, P183)
🧬 Was bedeutet es, wenn Marker anderer Haplogruppen auftauchen?
"Population Admixture: Over generations, populations can intermingle, leading to the mixing of haplogroups. For example, if a population with a distinct Y haplogroup interbreeds with another population, some descendants may carry Y chromosomes from both haplogroups, even if only one haplogroup is passed to male offspring." (quora.com/How-can-Haplogroups-mix[...], 2024).
copilot.microsoft.com: "Wenn bei einem Mann, der eindeutig zur Haplogruppe I1a2a1a1a1 gehört, Marker auftauchen, die typischerweise mit anderen Linien wie J2a (L152) oder E1b1a1a1 (L88, P-269) assoziiert sind, gibt es mehrere mögliche Erklärungen:
1. Technische Artefakte oder Testrauschen
- Manche SNPs (Single Nucleotide Polymorphisms) sind nicht exklusiv für eine Haplogruppe, sondern können konvergent auftreten - also unabhängig voneinander in verschiedenen Linien entstehen.
- Besonders bei sehr tiefen Subclades oder seltenen Mutationen kann es zu Fehlzuordnungen kommen, wenn ein Marker zufällig auch in einer anderen Linie mutiert ist.
2. Alte geteilte Ursprünge
- Haplogruppen wie I, J und E gehören alle zur größeren CT-M168-Linie. In sehr frühen Phasen der Menschheitsgeschichte (vor 40.000-60.000 Jahren) könnten bestimmte Marker gemeinsam vererbt worden sein, bevor sich die Linien trennten.
- Solche Marker sind dann prähistorisch alt und nicht mehr aussagekräftig für moderne Zuordnungen.
3. Testdesign oder Plattformunterschiede
- Manche kommerzielle Tests (z. B. YSEQ, FTDNA, Nebula) testen Marker, die nicht exklusiv für eine Linie sind, sondern eher zur Erkennung von Mutationsmustern dienen.
- Es kann sein, dass ein Marker wie L152 (typisch für J2a) auch in anderen Linien mutiert ist, aber dort keine phylogenetische Bedeutung hat.
4. Seltene oder unbekannte Rekombinationen?
- Das Y-Chromosom rekombiniert kaum - aber Pseudogenregionen oder Palindrombereiche können gelegentlich zu technischen Kreuztreffern führen.
- In sehr seltenen Fällen könnten auch Laborfehler oder Kontaminationen eine Rolle spielen.
Fazit
Wenn du I1a2a1a1a1 bist, dann ist das deine klare väterliche Linie. Das Auftreten einzelner Marker aus J2a oder E1b1a1a1 bedeutet nicht, dass du genetisch 'gemischt' bist - sondern eher, dass diese Marker entweder unspezifisch, konvergent oder technisch erklärbar sind".
"Haplogroup F, also known as F-M89 and previously as Haplogroup FT, is a very common Y-chromosome haplogroup. The clade and its subclades constitute over 95% of paternal lineages outside of Africa", "Possible time of origin 57,500-62,500 (Raghavan 2014), 45,000-55,700 BP (Karafet 2008), 43,000-56,800 BP (Hammer & Zegura 2002). Possible place of origin West or South Asia or Southeast Asia" (WP).
"One study, which did not comprehensively screen for other subclades of F-M89 (including some subclades of GHIJK), found that Indonesian men with the SNP P14/PF2704 (which is equivalent to M89), comprise 1.8% of men in West Timor, 1.5% of Flores 5.4% of Lembata 2.3% of Sulawesi and 0.2% in Sumatra [Chiaroni et al. 2009, Tumonggor et al. 2014]. F1 (P91), F2 (M427) and F3 (M481; previously F5) are all highly rare and virtually exclusive to regions/ethnic minorities in Sri Lanka, India, Nepal, South China, Thailand, Burma, and Vietnam" (WP).
"F-L352/PF2728/M3734" (genetiker.wordpress.com/more-y-snp-calls-for-an-early-neolithic-hungarian-genome, 2015, "negative calls are in non-bold").
"Haplogroup IJK is a human Y-chromosome DNA haplogroup. IJK is a primary branch of the macrohaplogroup HIJK. Its direct descendants are haplogroup IJ and haplogroup K", "Possible time of origin 49,000-59,000 BP. Possible place of origin Eurasia" (WP).
"Haplogroup IJ (M429/P125) is a human Y-chromosome DNA haplogroup, an immediate descendant of Haplogroup IJK (formerly known as Haplogroup F-L15). IJK is a branch of Haplogroup HIJK", "9,700-44,600 years BP. Possible place of origin South West Asia, Caucasus" (WP).
"Haplogroup I (M170) is a Y-chromosome DNA haplogroup. It is a subgroup of haplogroup IJ, which itself is a derivative of the haplogroup IJK", "Possible time of origin ~42,900 Years BP, Coalescence age ~27,500 Years BP. Possible place of origin Europe" (WP).
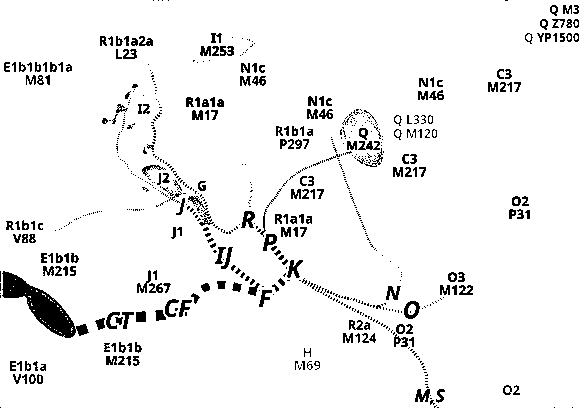
Abb. "Y-DNA haplogroup diversion spread. Overview of a possible evolution of the Y-chromosome haplogroups beginning with branch CT. The dominant haplogroups in the euroasiatic and nordeastafrican regions are shown", "Actual HG-Dominance by crosschecking maps of Eupedia, GeneticAtlas and Robertius (Wikipedia EU map) and other Wikipedia HG maps", Chris R. @AlpGen, June 2012, under Creative Commons Licence CC BY-SA 4.0.
Haplogroup I1 (M253: The Gravettian and the Vikings 3-5 milleniums ago, "[f]ound mainly in Northern Europe", WP)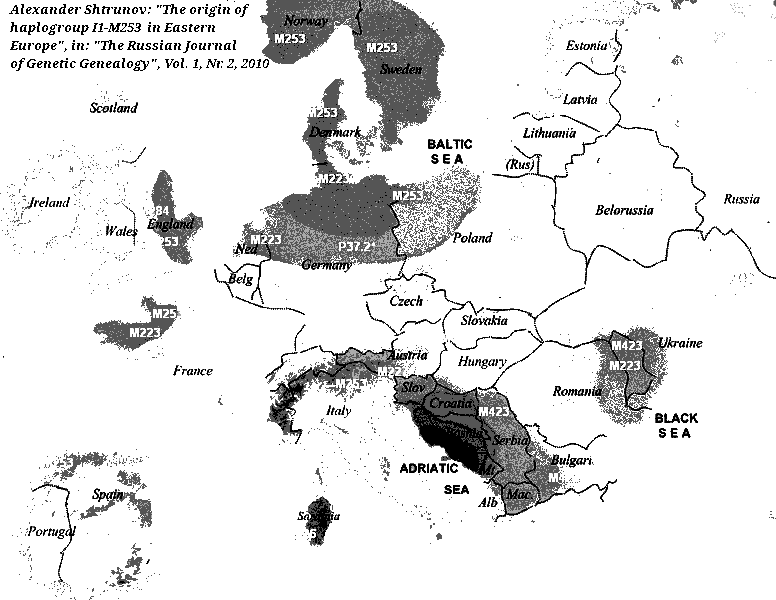
"Haplogroup I-M253, also known as I1, is a Y chromosome haplogroup", "Possible time of origin 3,170-4,600-5,070 BP (today's diversification), previously 11,000 BP to 33,000 BP, 27,500 (diversification with I2-FGC77992). Possible place of origin Northern Europe" (WP).
"Räumlich könnte die Y-Haplogruppe I in einem Refugium auf der Balkanhalbinsel bis zum Schwarzen Meer überdauert haben. Beides lässt sich am ehesten mit der Gravettien-Kultur verbinden. Mit dem Rückgang der Gletscher breiteten sich die Träger dieses Gens dann im Nordwesten Europas, vor allem in Skandinavien, aus"; "Höchste Frequenzen: Kroaten in Bosnien und Herzegowina 73,3 %, Bosniaken 58 %, Darginer 58 %, Kroaten 43,8-51 %, Schweden 44 %, Sarden 42,3 %, Bosnische Serben 31 %, Serben 31,5 %, Norweger 40,3 %, Deutsche 38 %, Dänen 36 %, Isländer 33 %, Kosovo-Albaner 30 %, Finnen 29 %, Mazedonier 25 %, Kurden 25 %, Niederländer 25 %, Engländer +20 %, Rumänen +20 %, Bulgaren +20 %"; "I1 (M253): Hohe Verbreitung in Skandinavien und dem nördlichen Mitteleuropa" (WP).
 "The Gravettian was an archaeological industry of the European Upper Paleolithic that succeeded the Aurignacian circa 33,000 years BP. It is archaeologically the last European culture many consider unified, and had mostly disappeared by c. 22,000 BP, close to the Last Glacial Maximum, although some elements lasted until c. 17,000 BP. In Spain and France, it was succeeded by the Solutrean and by the Epigravettian in Italy, the Balkans, Ukraine and Russia. The Gravettian culture is known for their artistic works including the famous Venus figurines, which were typically carved from either ivory or limestone"; "[d]er Begriff Gravettien wurde 1938 von Dorothy Garrod eingeführt, nach Funden im Abri La Gravette bei Bayac im Département Dordogne" (WP).
"The Gravettian was an archaeological industry of the European Upper Paleolithic that succeeded the Aurignacian circa 33,000 years BP. It is archaeologically the last European culture many consider unified, and had mostly disappeared by c. 22,000 BP, close to the Last Glacial Maximum, although some elements lasted until c. 17,000 BP. In Spain and France, it was succeeded by the Solutrean and by the Epigravettian in Italy, the Balkans, Ukraine and Russia. The Gravettian culture is known for their artistic works including the famous Venus figurines, which were typically carved from either ivory or limestone"; "[d]er Begriff Gravettien wurde 1938 von Dorothy Garrod eingeführt, nach Funden im Abri La Gravette bei Bayac im Département Dordogne" (WP).
"Während der Wikingerzeit erlebte I-M253 eine weitere Expansion. Margaryan et al. 2020 analysierten 442 Individuen aus der Wikingerwelt von verschiedenen archäologischen Stätten in Europa. I-M253 war die in der Studie am häufigsten gefundene Y-Haplogruppe [95 samples]. Norwegische und dänische Wikinger brachten mehr I1 nach Großbritannien und Irland, während schwedische Wikinger es nach Russland und in die Ukraine einführten und mehr davon nach Finnland und Estland brachten", zusammengefasst aber waren Ableitungen von R1 am häufigsten [siehe unten]: "R1b (84 samples) and R1a, especially (but not exclusively) of the Scandinavian R1a-Z284 subclade (61 samples)" (WP).
- I1 (M253, L495, L845/YSC0000278, isogg.org/tree/ISOGG_HapgrpI.html)
- I1a (M307, P30, P40, S62, S63, S64, S65, S66, DF29/S438 [M307+, M307.1+, M307.2+])
- I1a*
- I1a1 (M227, formerly I1a4)
- I1a1a (M72, formerly I1a3)
- I1a2 (M21, wikidoc.org/index.php/Haplogroup_I1a_(Y-DNA), Z58/S244)
- I1a2a (Z59/S246)
- I1a2a1~ (S337/Z60, S439/Z61)
- I1a2a1a (S440/Z140, Z141) [Z141 = Gothic / Wielbark]
- I1a2a1a1 (S1953/Z2535, isogg.org/tree/ISOGG_HapgrpI.html: "Mutations previously on tree but found so far only in one man or in closely related men: [...] M21 [...]")
- I1a2a1a (S440/Z140, Z141) [Z141 = Gothic / Wielbark]
- I1a2a1~ (S337/Z60, S439/Z61)
- I1a2a (Z59/S246)
- I1a (M307, P30, P40, S62, S63, S64, S65, S66, DF29/S438 [M307+, M307.1+, M307.2+])
 Abb./Fig. oben rechts Alexander Shtrunov: "The Origin of Haplogroup I1-M253 in Eastern Europe", in: "The Russian Journal of Genetic Genealogy", Vol. 1, Nr. 2, 2010 (with M253, M423, M223, M227, M26, M284, P37.2). Sjur Cappelen Papazian: "The origin of haplogroup I1-M253 in Eastern Europe", in: "Cradle of Civilization", 20. Nov. 2013: "In 11-10th millennium BC global climate changes took place, [...]. Roots of haplogroup I1 evidently came from such Paleolithic cultures as Ahrensburgian and Swiderian; its carriers represented were the part of autochthonous population of Northern and Eastern Europe. The main activities of carriers of haplogroup I1 were hunting and gathering. [...] Carriers of haplogroup I1 were speakers of Paleo-European language, which didn't belong tothe Uralic or Indo-European families. Its traces were reveiled in the European toponomy and in the Sami language" (aratta.wordpress.com/2013/11/28/the-origin-of-haplogroup-i1-m253-in-eastern-europe). Lázló Pál, Budapest: "Until now, no strong correlation was found between the haplogroup I and existing languages/language families. But we know that people with haplogroup I populated huge areas in Europe before the Uralic and the subsequent Indo-European migration, and their proportion is still high in Nothern Europe, on the Balkan and among Uralic speakers as well. Studying the substratum effects of the relevant languages can be the only method to know more about their (probably long ago extinct) language(s). The best source can be the German / Germanic, a considerable proportion of its basic vocabulary can not be derived from PIE, and according to some linguists, even 50% of the German irregular verbs (these are important building blocks of the German language) can not be derived from Proto-Indo-European roots. (And cannot be derived from Uralic, Basque etc). Perhaps these are the only known traces of this mysterious I-language" (quora.com, 2020).
Abb./Fig. oben rechts Alexander Shtrunov: "The Origin of Haplogroup I1-M253 in Eastern Europe", in: "The Russian Journal of Genetic Genealogy", Vol. 1, Nr. 2, 2010 (with M253, M423, M223, M227, M26, M284, P37.2). Sjur Cappelen Papazian: "The origin of haplogroup I1-M253 in Eastern Europe", in: "Cradle of Civilization", 20. Nov. 2013: "In 11-10th millennium BC global climate changes took place, [...]. Roots of haplogroup I1 evidently came from such Paleolithic cultures as Ahrensburgian and Swiderian; its carriers represented were the part of autochthonous population of Northern and Eastern Europe. The main activities of carriers of haplogroup I1 were hunting and gathering. [...] Carriers of haplogroup I1 were speakers of Paleo-European language, which didn't belong tothe Uralic or Indo-European families. Its traces were reveiled in the European toponomy and in the Sami language" (aratta.wordpress.com/2013/11/28/the-origin-of-haplogroup-i1-m253-in-eastern-europe). Lázló Pál, Budapest: "Until now, no strong correlation was found between the haplogroup I and existing languages/language families. But we know that people with haplogroup I populated huge areas in Europe before the Uralic and the subsequent Indo-European migration, and their proportion is still high in Nothern Europe, on the Balkan and among Uralic speakers as well. Studying the substratum effects of the relevant languages can be the only method to know more about their (probably long ago extinct) language(s). The best source can be the German / Germanic, a considerable proportion of its basic vocabulary can not be derived from PIE, and according to some linguists, even 50% of the German irregular verbs (these are important building blocks of the German language) can not be derived from Proto-Indo-European roots. (And cannot be derived from Uralic, Basque etc). Perhaps these are the only known traces of this mysterious I-language" (quora.com, 2020).
Abb./Fig. links "Vénus de Lespugue", "mammoth ivory, found in the Grotte des Rideaux (Lespugue, Haute-Garonne, France). Dated from the Gravettian, Upper Paleolithic, 23 000 years. Currently in the Musée de l'Homme, in Paris", 2018, von Vassil, gemeinfrei (modifiziert).
Haplogroup I2 (Early Neolithic "Nuragic" Admixtures: From Persia to Italy, Sardinia, Spain)
"I-P222 (P222, PF3861, S118, U250, rs17315723). Found in haplogroup I2a1b1a on FTDNA tree. Coincident with Y3259 (Sims et al 2007). Ancestral: C. Derived: G. Chrom.: Y. Pos.: 16776320 (hg38) - 18888200 (hg19)" (genetichomeland.com/welcome/dnamarkerindex.asp).
- "M170 [Haplogroup I, "is found mainly in Europe and the Caucasus", WP]
- P215 (M438, PF3853, S31, I2, rs17307294). Defines major haplogroup I2 on ISOGG and FTDNA trees. Arose about 26,000 bce ["[f]ound mainly in Balkans, Southeast Europe and Sardinia", WP].
 CTS2257 (PF3704, Z2647, rs753852546). Defines haplogroup I2a on FTDNA tree. Believed coincident with CTS1799 and Z2672 ("The great winner during the Neolithic period [ca. 7000-1700 BCE] was haplogroup I2a, which consistently shows up alongside G2a [L140, M406] in most Neolithic sites tested to date (Starčevo, Körös, Lengyel, LBK, Cardium Pottery, Megalithic), and seem to increase in frequency over time and as one moves towards Northwest Europe. Based on the few samples available it appears to have been particularly common in the Megalithic culture. All four Megalithic Y-DNA samples (from France and Spain) belonged to I2a1 or I2a2. [...] Therefore, although I2a was just one of many Mesolithic hunter-gatherers' lineages in Europe when agriculturists arrived, it is the only one that readily embraced the new lifestyle and managed to supersede the original farmers in number. I2a's destiny was not only linked to its ability to chum with G2a, but we could say that G2a farmers catalysed I2a's success. I2a people integrated G2a tribes, learned the new Neolithic techniques from them and became so good at them that over time the student overtook the master", eupedia.com/forum/threads/the-great-pairings-of-y-dna-haplogroups-in-prehistory.31431, Maciamo, 19. Jul. 2015)
CTS2257 (PF3704, Z2647, rs753852546). Defines haplogroup I2a on FTDNA tree. Believed coincident with CTS1799 and Z2672 ("The great winner during the Neolithic period [ca. 7000-1700 BCE] was haplogroup I2a, which consistently shows up alongside G2a [L140, M406] in most Neolithic sites tested to date (Starčevo, Körös, Lengyel, LBK, Cardium Pottery, Megalithic), and seem to increase in frequency over time and as one moves towards Northwest Europe. Based on the few samples available it appears to have been particularly common in the Megalithic culture. All four Megalithic Y-DNA samples (from France and Spain) belonged to I2a1 or I2a2. [...] Therefore, although I2a was just one of many Mesolithic hunter-gatherers' lineages in Europe when agriculturists arrived, it is the only one that readily embraced the new lifestyle and managed to supersede the original farmers in number. I2a's destiny was not only linked to its ability to chum with G2a, but we could say that G2a farmers catalysed I2a's success. I2a people integrated G2a tribes, learned the new Neolithic techniques from them and became so good at them that over time the student overtook the master", eupedia.com/forum/threads/the-great-pairings-of-y-dna-haplogroups-in-prehistory.31431, Maciamo, 19. Jul. 2015)
Fig./Abb. "Megalith-Reihen bei Carnac, Bretagne, Frankreich", 2002, by Rainer Zenz / Brudersohn, under Creative Commons Licence CC BY-SA 3.0 (modified).
- L460 (PF3647, S238 rs7892855). Defines haplogroup I2a1 on ISOGG, YFull, and FTDNA trees.
- I2a1a (M26, P.37.2)
- I2a1a1 (L160,"Dominante Haplogruppe in Sardinien (Gründereffekt), auch in niedriger Frequenz in den Pyrenäen, der gesamten Atlantikküste und auf Inseln des Atlantiks von Schottland/Norwegen bis zur Sahara/Kanaren gefunden", WP)
- I2a1a1a (L672/S327, ISOGG build 11.128, "[PMID 18385274] This snp distinguishes haplogroups", snpedia.com/index.php/Rs112989530, "I2a1a1a (L672) was already found in Mesolithic Sweden, which implies that I2a1a had a very wide distribution from Iberia to Scandinavia during the Mesolithic period. Later, they would have adopted agriculture by intermixing with Near Eastern newcomers", eupedia.com/europe/Haplogroup_I2_Y-DNA.shtml, "[a] genetic study published in Science in March 2019 [Olalde et al.] found a significant increase in north-central European ancestry in Iberia during the transition from the Bronze Age [3300-1200 BC] to the Iron Age. The authors of the study suggested that the spread of the Urnfield culture was associated with this transition, during which the Celtiberians may have emerged. A Celtiberian male examined in the study was found to be a carrier of the paternal haplogroup I2a1a1a", WP; I2a1a1a L158, M26, ISOGG 2019, "Sardinians")
- I2a1a2 (M423, Grasgruber et al.: "Mapping the Mountains of Giants. Anthropometric Data from the Western Balkans Reveal a Nucleus of Extraordinary Physical Stature in Europe", in: Biology", Vol. 11, 2022, p. 786; L178, "I2a1b (M423, L178) was known as I1b until 2007, and I2a2 from 2008 to 2010", eupedia.com)
- I2a1a2b ("L621 is typical of the Slavic populations, being highest in Southeastern European regions of Bosnia-Herzegovina and South Croatia (>45%), in Croats (37.7-69.8%), Bosniaks (43.53-52.17%), and Serbs (36.6-42%) - often called 'Dinaric'", WP).
- I2a1a1 (L160,"Dominante Haplogruppe in Sardinien (Gründereffekt), auch in niedriger Frequenz in den Pyrenäen, der gesamten Atlantikküste und auf Inseln des Atlantiks von Schottland/Norwegen bis zur Sahara/Kanaren gefunden", WP)
- P214 (M436, PF3856, S33 rs17315680). Defines haplogroup I2a1b on ISOGG, YFull, and FTDNA trees. Often referred to as haplogroup I2a2 in old literature. Dispersed throughout Mesolithic Europe (for an alternative classification as I2b see above).
 M223 (rs367573274, I2a1b1). A major branch of haplogroup I2a. Formerly labeled haplogroup I2d in the literature. Example is ancient sample R15 from central Italy circa 7125 bce. Originally enumerated as a C->T mutation allele ("A genetic study published in Nature Communications in February 2020 [Marcus et al. 2020] examined the remains of 17 individuals identified with Nuragic civization [on Sardinia]. The samples of Y-DNA extracted belonged to haplogroup I2a1b1 ([M223], 2 samples), R1b1b2a [L23/S141, L49], G2a2b2b1a1 [PF3433, PF2829, PF3349], R1b1b ([L754], 4 samples), J2b2a1 ([L283], 3 samples) and G2a2b2b1a1a", WP).
M223 (rs367573274, I2a1b1). A major branch of haplogroup I2a. Formerly labeled haplogroup I2d in the literature. Example is ancient sample R15 from central Italy circa 7125 bce. Originally enumerated as a C->T mutation allele ("A genetic study published in Nature Communications in February 2020 [Marcus et al. 2020] examined the remains of 17 individuals identified with Nuragic civization [on Sardinia]. The samples of Y-DNA extracted belonged to haplogroup I2a1b1 ([M223], 2 samples), R1b1b2a [L23/S141, L49], G2a2b2b1a1 [PF3433, PF2829, PF3349], R1b1b ([L754], 4 samples), J2b2a1 ([L283], 3 samples) and G2a2b2b1a1a", WP).
Fig./Abb. "Bronzetti nuragici", Museo archeologico nazionale G.A. Sanna - MIBAC, 2012, by Cristiano Cani under Creative Commons Licence CC BY-SA 3.0 (modified).
- Y3259 (FGC3552, rs758826454). On YFull tree as an intermediate branch under M223. Believed coincident with P222. Marker [P222] currently considered coincident with marker [Y3259], using that phylogenetic tree" (genetichomeland.com/welcome/dnapedigree.asp?RecordID=33154).
- Y3259 (FGC3552, rs758826454). On YFull tree as an intermediate branch under M223. Believed coincident with P222. Marker [P222] currently considered coincident with marker [Y3259], using that phylogenetic tree" (genetichomeland.com/welcome/dnapedigree.asp?RecordID=33154).
- I2a1a (M26, P.37.2)
- L460 (PF3647, S238 rs7892855). Defines haplogroup I2a1 on ISOGG, YFull, and FTDNA trees.
- "I2b (M436/P214/S33, P216/S30, P217/S23, P218/S32) (formerly I1b2, alternatively I2a1b)
- I2b1 (M223, P219/S24, P220/S119, P221/S120, P222/U250/S118, P223/S117, formerly I1b2a - old I1c, "[o]ccurs at a moderate frequency among populations of Northwest Europe, with a peak frequency in the region of Lower Saxony in central Germany; minor offshoots appear in Moldavia and Russia (especially around Vladimir, Ryazan, Nizhny Novgorod, and the Republic of Mordovia), and among speakers of Persian (including Iranians, Hazaras in Afghanistan and Pakistan, and Tajiks in Afghanistan)", WP; "I2b1 dates to 8000 or 9000 BC. It appears to have originated in the German Rhineland. Today it is much more common in the German Rhineland and in the Netherlands than anyone else. The notion that M284 developed in Britain around 2000 BC, but possibly not long after I2b1 itself developed, supports thinking that the direct ancestors of M284 may have reached Great Britain not long after the last ice age", sites.rootsweb.com/~villandra/McKinstry/DeepHistory.html).
- M284 (formerly I1b2a1, "a typical British subclade", "I2b1a (M284) has been found almost exclusively among the population of Great Britain, suggesting that the clade may have arisen in that island. I2b1a is comparatively rare in Ireland. Where it is found in those of Irish descent with Gaelic surnames, and particularly in baronial families with a credible pedigree back to a Cruithin (British) origin, this suggests an ancestor who arrived in Ireland from Celtic Britain. For example it is found in McGuinness and McCartan men descended from the Uí Echach Cobha, a lineage considered Cruithin in the 6th century AD", geni.com/projects/I-M284-Y-DNA/3784, 2009)
- M379 (formerly I1b2a2)
- P78 (formerly I1b2a3)
- P95 (formerly I1b2a4)
- I2b2 (L38/S154, L39/S155, L40/S156, L65/S159, isogg.org/tree/2008/ISOGG_HapgrpI08.html)
- I2b1 (M223, P219/S24, P220/S119, P221/S120, P222/U250/S118, P223/S117, formerly I1b2a - old I1c, "[o]ccurs at a moderate frequency among populations of Northwest Europe, with a peak frequency in the region of Lower Saxony in central Germany; minor offshoots appear in Moldavia and Russia (especially around Vladimir, Ryazan, Nizhny Novgorod, and the Republic of Mordovia), and among speakers of Persian (including Iranians, Hazaras in Afghanistan and Pakistan, and Tajiks in Afghanistan)", WP; "I2b1 dates to 8000 or 9000 BC. It appears to have originated in the German Rhineland. Today it is much more common in the German Rhineland and in the Netherlands than anyone else. The notion that M284 developed in Britain around 2000 BC, but possibly not long after I2b1 itself developed, supports thinking that the direct ancestors of M284 may have reached Great Britain not long after the last ice age", sites.rootsweb.com/~villandra/McKinstry/DeepHistory.html).
- P215 (M438, PF3853, S31, I2, rs17307294). Defines major haplogroup I2 on ISOGG and FTDNA trees. Arose about 26,000 bce ["[f]ound mainly in Balkans, Southeast Europe and Sardinia", WP].
"Haplogroup J-M304, also known as J is a human Y-chromosome DNA haplogroup. It is believed to have evolved in Western Asia. The clade spread from there during the Neolithic, primarily into North Africa, the Horn of Africa, the Socotra Archipelago, the Caucasus, Europe, Anatolia, Central Asia, South Asia, and Southeast Asia" (WP).
- J (12f2.1, M304/PAGES00016/PF4609, S6, S34, S35)
- J1 ("Semitid/Bedouinid Arabids (M267)", WP)
- J1a (M62)
- J1b (M365)
- J1c (M367, M368)
- J1d (M369)
- J1e (M390)
- J2 (M172, PAGES00028, L228, "Syrid/Nahrainid Arabids (M172) [...] Anatolia, Greece, the Balkans, Italy, Iran, the Caucasus, South Asia, and Central Asia", WP, "Haplogroup J2 is said to have originated in the Levant, Mesopotamia or Iran about 32 000 years ago", quora.com, 2023)
- J2a (M410, L152, L212/PF4988, L559/PF4986, "J2a spreads from Europe to Southern Asia, with presence also on the African Mediterranean coast. The center of distribution and probable origin is in Northern Fertile Crescent between Asia Minor (modern eastern Turkey), Syria, Armenia, Caucasus and Mesopotamia (modern Iraq)", isogg.org/wiki/Haplogroup_J2a_(Y-DNA))
- J2a*
- J2a1 (DYS413, isogg.org; CTS3547, CTS9718/PF5102, L26/S57/PF5110/PAGES00055, +28 SNPs, formed 18300 ybp, TMRCA 16500 ybp, yfull.com/arch-3.09/tree/J2a1)
- J2a1i ("J2a1i L88.2, L198", isogg.org/tree/2012/ISOGG_HapgrpJ12.html, [Proband A: "J2-L88 (J2a1i) [...] My paternal side is of Galician Sephardic origin (Ardon, Spain), but I have not heard of anyone else with similar heritage and same haplogroup", genoplot.com, Proband B: "I'm Palestinian my haplogroup r-L23 [=R1b1a2a, depend. from M343, P297, M269] ([Z2019]?), What's next?", "[...] M96, P222/U250/S118, L88, L88.1 [...]", 2018])
- J2a2 (M340)
- J2a1 (DYS413, isogg.org; CTS3547, CTS9718/PF5102, L26/S57/PF5110/PAGES00055, +28 SNPs, formed 18300 ybp, TMRCA 16500 ybp, yfull.com/arch-3.09/tree/J2a1)
- J2a*
- J2b (M12, M314, M221; also M102, Z2497, Z2499 Z2500/CTS10660)
- J2b* (At least one living example found in Uzbekistan)
- J2b1 (M205; includes former J2b1b/J-PF7344)
- J2b1* (Found in ancient remains (3,650 years BP) in Lebanon)
- J2b1a (YP31, Y3165, Y167062) Living examples found in Greece and Italy" (WP).
- J2b2 (Z1297)
- J2b2a (M241, YFull 9700 ybp)
- J2b2a1 ("Archäogenetische Studien haben gezeigt, dass sich eine wichtige Y-DNA-Haplogruppe unter den Illyrern, J2b-L283 [J2b2a1], über die Cetina-Kultur über die östliche Adria vom Cetina-Tal in Kroatien bis nach Montenegro und Nordalbanien verbreitete", WP).
- J2b2a (M241, YFull 9700 ybp)
- J2b1 (M205; includes former J2b1b/J-PF7344)
- J2b* (At least one living example found in Uzbekistan)
- J2a (M410, L152, L212/PF4988, L559/PF4986, "J2a spreads from Europe to Southern Asia, with presence also on the African Mediterranean coast. The center of distribution and probable origin is in Northern Fertile Crescent between Asia Minor (modern eastern Turkey), Syria, Armenia, Caucasus and Mesopotamia (modern Iraq)", isogg.org/wiki/Haplogroup_J2a_(Y-DNA))
- J1 ("Semitid/Bedouinid Arabids (M267)", WP)
Markers from E1: Subclads E1b1a1a1 (L88/L88.1), E1b1a1a1h (P-269)
"E1b1a1a1h is defined by markers P268 and P269. It was first reported in a person from the Gambia (Karafet / Mendez et al. 2008 ['New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree', in: 'Genome Research', Vol. 18, Nr. 5, May 2008, p. 830-838, pmc.ncbi.nlm.nih.gov/articles/PMC2336805]"; "African admixture in Europe refers to the presence of human genotypes attributable to periods of human population dispersals out of Africa in the genetic history of Europe. More recent African admixture - primarily Berber admixture from North Africa - is associated with historic migrations through the Mediterranean Sea and the Muslim conquests of the Early Middle Ages. This admixture can be found primarily in the Iberian Peninsula (modern day Spain and Portugal), with higher levels in the West and the South and Southern Italy, with higher levels in Sardinia and Sicily" (WP).
- CT
- DE (M1/YAP, M145/P205, M203, P144, P153, P165, P167, P183)
- E (M96)
- E1 (P147)
- E1b (P177)
- E1b1 (P2)
- E1b1a (V38)
- E1b1a1 (DYS271/M2/SY81, M291, P1/PN1, P189, P293, V43, V95, Z1101, Z1107, Z1116, Z1120, Z1122, Z1123, Z1124, Z1125, Z1127, Z1130, Z1133)
- E1b1a1a (L576)
- E1b1a1a1 (L86.1, L88.3, M180/P88, PAGES00066, P182, Z1111, Z1112, "E1b1a1a1-L88.2/L88.3/L431/M4851/L88.1/L88", genetiker.wordpress.com/y-snp-calls-for-i1532", 2015)
- E1b1a1a1h (P268, P269)
- E1b1a1a1 (L86.1, L88.3, M180/P88, PAGES00066, P182, Z1111, Z1112, "E1b1a1a1-L88.2/L88.3/L431/M4851/L88.1/L88", genetiker.wordpress.com/y-snp-calls-for-i1532", 2015)
- E1b1a1a (L576)
- E1b1a1 (DYS271/M2/SY81, M291, P1/PN1, P189, P293, V43, V95, Z1101, Z1107, Z1116, Z1120, Z1122, Z1123, Z1124, Z1125, Z1127, Z1130, Z1133)
- E1b1a (V38)
- E1b1 (P2)
- E1b (P177)
- E1 (P147)
- E (M96)
- DE (M1/YAP, M145/P205, M203, P144, P153, P165, P167, P183)
"Haplogroup E-M2, also known as E1b1a1-M2, is a human Y-chromosome DNA haplogroup. E-M2 is primarily distributed within Africa followed by West Asia. More specifically, E-M2 is the predominant subclade in West Africa, Central Africa, Southern Africa, and the region of the African Great Lakes; it also occurs at moderate frequencies in North Africa, and the Middle East. E-M2 has several subclades, but many of these subhaplogroups are included in either E-L485 or E-U175"; "Possible time of origin 39,200 years BP. Coalescence age 16,300 years BP. Possible place of origin West Africa or Central Africa" (WP).
 "E1b1a1a1 [...] formed 15000 ybp, TMRCA 10600 ybp" (yfull.com/arch-3.10/tree/E1b1a1a1). Diane Rowold et al.: "At the southeast fringe of the Bantu expansion: genetic diversity and phylogenetic relationships to other sub-Saharan tribes", in: "Meta Gene", 2. Oct. 2014, Vol. 2, S. 670-685: "As indicated by the Y-SNP haplogroup frequencies (Table 2), the E1b1a1a1-M180, and B2a1a-M109 mutations, both of which are signatures of the Bantu expansion, rank as the two most abundant Y-SNP haplogroups of the MAP [Maputo Province Bantu] population (at 71.8% and 14.1%, respectively). These haplogroups are followed in frequency by the Bantu and Eurasian markers E2b (7.7%) that include E2b-M54, E2b1-M85 and E2b1a-M200, and R1a1a-M198 (2.6%), respectively. The ancient East African haplogroup A1b1b2b1-M118 [M118+, M118-, M118-] and the Pygmy haplogroup B2b-M112 [M112-, M112-] are represented by one (1.3%) individual each." (pmc.ncbi.nlm.nih.gov/articles/PMC4287857).
"E1b1a1a1 [...] formed 15000 ybp, TMRCA 10600 ybp" (yfull.com/arch-3.10/tree/E1b1a1a1). Diane Rowold et al.: "At the southeast fringe of the Bantu expansion: genetic diversity and phylogenetic relationships to other sub-Saharan tribes", in: "Meta Gene", 2. Oct. 2014, Vol. 2, S. 670-685: "As indicated by the Y-SNP haplogroup frequencies (Table 2), the E1b1a1a1-M180, and B2a1a-M109 mutations, both of which are signatures of the Bantu expansion, rank as the two most abundant Y-SNP haplogroups of the MAP [Maputo Province Bantu] population (at 71.8% and 14.1%, respectively). These haplogroups are followed in frequency by the Bantu and Eurasian markers E2b (7.7%) that include E2b-M54, E2b1-M85 and E2b1a-M200, and R1a1a-M198 (2.6%), respectively. The ancient East African haplogroup A1b1b2b1-M118 [M118+, M118-, M118-] and the Pygmy haplogroup B2b-M112 [M112-, M112-] are represented by one (1.3%) individual each." (pmc.ncbi.nlm.nih.gov/articles/PMC4287857).
"Bantu Expansion": "Chronological overview after Nurse and Philippson (2003):
1 = 4,000-3,500 BP: origin [highlands between Cameroon and Nigeria, Mambilla region],
2 = 3,500 BP: initial expansion ('early split': 2.a = Eastern [e.g. Democratic Republic of Congo], 2.b = Western [e.g. Congo, Gabun])
3 = 2,000-1,500 BP: Urewe nucleus of Eastern Bantu
4-7: southward advance
8 = 2,500 BP: Congo nucleus
9 = 2,000-1,000 BP: last phase" (WP).
Abb./Fig. Westafrica, openstreetmap.de/karte, Database Contents License: opendatacommons.org/licenses/dbcl/1.0.
Nogeiro et al.: "Portuguese crypto-Jews: the genetic heritage of a complex history", in: "Frontiers in Genetics", Nr. 6, Feb. 2015, listet in unterschiedlicher Sample-Verteilung für fünf Standorte (Marocco, Mediterrean, Libya, Bulgaria, Turkey): E-M2, E-M35, E-M78, E-M123, G-M201, I-M170 (only Bulgaria), J-12f2.1, J2-M172, KLT-M9, Q-M242, R-M207, R-M17, R-P25, R-M269.
1900-950 BCE: I1 related tree for latest marker S1954/YSC0000261 (Z140/S440, S1953/Z2535 → I1a2a1a1a1a)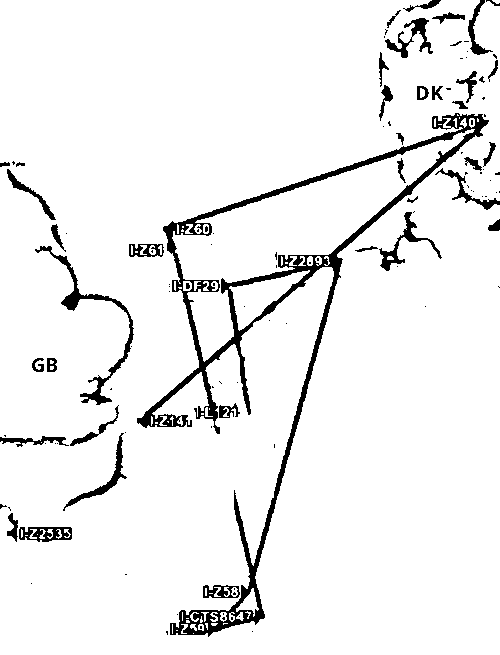
"The I-DF29 paternal line [I1a] was formed when it branched off from the ancestor I-M253 and the rest of humankind around 2600 BCE. He is the ancestor of at least 11 descendant lineages known as I-Z63, I-Z58, I-FGC15560, I-A5716, I-Y14628, I-Y18697, I-Y11204, I-Y2592, I-PH525, I-BY169148, & I-BY183623. There are 59.849 DNA tested descendants, and they specified that their earliest known origins are from: Sweden, United States, England and 121 other countries" (discover.familytreedna.com/y-dna/I-DF29/story).
Abb. phylogeographer.com/snp-lookup/?I-YSC261, phylogeographer.com/mygrations?hg=I1&clade=I-YSC261. Marker I-L21 (I1, 4.500 BC, Belgien), I-DF29, I-Z2893, I-Z58, I-Z59, I-CTS8647 (2.500 BC, Frankreich), I-Z60, I-Z61, I-Z140 (Dänemark), I-Z141, I-Z2535 (2.000 BC, Bretagne), I-YSC261 (1.500 BC). Leaflet, Map data: OpenStreetMap contributors, CC-BY-SA, Imagery: Mapbox.
"The I-Z58 paternal line [I1a2] was formed when it branched off from the ancestor I-DF29 and the rest of humankind around 2450 BCE. [...] He is the ancestor of at least 2 descendant lineages known as I-Z59 & I-Z138. There are 24.851 DNA tested descendants, and they specified that their earliest known origins are from: Sweden, United States, England and 92 other countries" (discover.familytreedna.com/y-dna/I-Z58/story).
"The man who is the most recent common ancestor of this line [Z59, I1a2a] is estimated to have been born around 2350 BCE. He is the ancestor of at least 5 descendant lineages known as I-CTS8647, I-Z2041, I-A12798, I-Y129360, & I-BY157528. There are 19.883 DNA tested descendants, and they specified that their earliest known origins are from: Sweden, United States, England and 84 other countries" (discover.familytreedna.com/y-dna/I-Z59/story).
"The I-CTS8647 paternal line [I1a2a-1] was formed when it branched off from the ancestor I-Z59 and the rest of humankind around 2350 BCE. He is the ancestor of at least 3 descendant lineages known as I-A11141, I-Z61, & I-FTD84484. There are 15.831 DNA tested descendants, and they specified that their earliest known origins are from: Sweden, United States, England and 72 other contries" (discover.familytreedna.com/y-dna/I-CTS8647/story).
"The man who is the most recent common ancestor of this line [Z61, I1a2a1] is estimated to have been born around 2200 BCE. He is the ancestor of at least 2 descendant lineages known as I-Z60 & I-S9939. There are 15.556 DNA tested descendants, and they specified that their earliest known origins are from: Sweden, United States, England and 71 other contries" (discover.familytreedna.com/y-dna/I-Z61/story).
"The I-Z60 paternal line was formed when it branched off from the ancestor I-Z61 and the rest of humankind around 2200 BCE. The man who is the most recent common ancestor of this line is estimated to have been born around 2100 BCE. He is the ancestor of at least 10 descendant lineages known as I-CTS7362, I-FGC23806, I-Z140, I-A9290, I-BY56697, I-A9103, I-FT220000, I-BY70880 and 2 yet unnamed lineages" (discover.familytreedna.com/y-dna/I-Z60/story). "The I-Z140 paternal line [I1a2a1a] was formed when it branched off from the ancestor I-Z60 and the rest of humankind around 2100 BCE. The man who is the most recent common ancestor of this line is estimated to have been born around 2000 BCE. He is the ancestor of at least 4 descendant lineages known as I-Z141, I-A16593, I-E564, & I-FT383503. There are 8.721 DNA tested descendants, and they specified that their earliest known origins are from: England, United States, Germany and 65 other countries" (discover.familytreedna.com/y-dna/I-Z140/story).
"1a2a1a (Z140): Wird vor allem in Deutschland und in geringerer Häufigkeit auch in Frankreich, den BeNeLux-Staaten und den britischen Inseln gefunden" (WP).
"I1-Z140, Z14/S440 comes off the main I1-M253 Z58 -> Z62 branch. It then divides into five main branches, Y6231+, A196+, A1374+, F2642+ and Z2535+ [...]. Z2535 includes subgroups defined by SNPs: L338, YSC261, S12289; Y8333, Y8334, S1990, S2001, Y4015; A375, S1972, A376, S2000, Y3166, A264, A681, A261" (wikitree.com/wiki/Category:Y-DNA_Haplogroup_I-Z140).
"The I-Z141 paternal line was formed when it branched off from the ancestor I-Z140 and the rest of humankind around 2000 BCE. The man who is the most recent common ancestor of this line is estimated to have been born around 1900 BCE. He is the ancestor of at least 11 descendant lineages known as I-Z2535, I-FGC22406, I-Y5497, I-CTS6739, I-P259, I-FT214006, I-BY476, I-FT216827, I-FT137449, I-FGC94381, & I-FTA48749" (discover.familytreedna.com/y-dna/I-Z141/story). "The I-S1953/Z2535 paternal line [I1a2a1a1, and I1a2a1a-1, S1954/YSC0000261] was formed when it branched off from the ancestor I-Z141 and the rest of humankind around 1900 BCE. The man who is the most recent common ancestor of this line is estimated to have been born around 1850 BCE. He is the ancestor of at least 2 descendant lineages known as I-L338 & I-CTS10937. There are 3.808 DNA tested descendants, and they specified that their earliest known origins are from: England, United States, Germany and 49 other countries" (discover.familytreedna.com/y-dna/I-Z2535/story).
"The I-L338 paternal line [I1a2a1a1a] was formed when it branched off from the ancestor I-Z2535 and the rest of humankind around 1850 BCE. The man who is the most recent common ancestor of this line is estimated to have been born around 950 BCE. He is the ancestor of at least 2 descendant lineages known as I-S12289 & I-FT456763. There are 3.106 DNA tested descendants, and they specified that their earliest known origins are from: England, United States, Scotland and 39 other countries" (discover.familytreedna.com/y-dna/I-L338/story).
"If you check the ISOGG 2014 Tree you will find L41/PF3787 [I1a2a1a1a-1] as a defining SNP for Hapogroup I at the highest level. This is an error in the FTDNA I Haplotree that Morley did not correct" (forums.familytreedna.com/forum/y-dna-haplogroup-project-forums/i/i1-dna-project, 2014).
- I1a2a1a1a1 S1953/Z2535 [formed 4100 ybp, TMRCA 4100 ybp, yfull.com/tree/I-Z2535, All Western World]
- I1a2a1a1a1a S1954/YSC0000261, S1962^/Z4731^, S1968/Z4730, S1969, S1978^/Z4736^, S1987/Z4729, Z4727, Z4728
- I1a2a1a1a1a1 L338/S197
- I1a2a1a1a1a1a~ A1944/Y15155
- I1a2a1a1a1a1a1~ A2398, A2401, A2404, A2407
- I1a2a1a1a1a1b~ A1940/Y12660, A1953/Y12330, A1967/Y12331, A1976/Y12661, A1991/Y12669, A2006/Y12333, A2007/Y12665, A2028/Y12667, A2051/Y12668, A2055/Y12669, A2059/Y12670, A2107/Y12666, A5454/Y12664
- I1a2a1a1a1a1b1~ A1972/Y12663, A1990/Y12662, A1996/Y12332
- I1a2a1a1a1a1b2~ A8666, A8670, A8671, A8672, A8673, A8675, A8677, Y29607
- I1a2a1a1a1a1b2a~ A8674, A8676, A8679
- I1a2a1a1a1a1a~ A1944/Y15155
- I1a2a1a1a1a2~ S12289
- I1a2a1a1a1a2a~ A1818/Y8335, A1830/Y14670, BY461/Y8333, BY462/Y8844
- I1a2a1a1a1a2a1~ BY463/Y8334, BY464/Y8336, BY465/Y8337, BY467/Y8845, BY468/Y8846, Y14669, Y28952
- I1a2a1a1a1a2a1a~ A4637
- I1a2a1a1a1a2a1a1~ Z11619, Z44259, Z44260
- I1a2a1a1a1a2a1a~ A4637
- I1a2a1a1a1a2a1~ BY463/Y8334, BY464/Y8336, BY465/Y8337, BY467/Y8845, BY468/Y8846, Y14669, Y28952
- I1a2a1a1a1a2b~ FGC13287/Y4015, S1990, S2001
- I1a2a1a1a1a2b1~ CTS6688, CTS8611, Z44256, Z44257, Z44258
- I1a2a1a1a1a2b2~ A1627, A1631
- I1a2a1a1a1a2b3~ A8601
- I1a2a1a1a1a2b3a~ A8596, A8600, A8604, A8605, A8606
- I1a2a1a1a1a2b4~ A375/FGC13289, S1965, S1975, S1977, S2014
- I1a2a1a1a1a2b4a~ A1790/FGC36457, A1791, A1794/FGC36550
- I1a2a1a1a1a2b4b~ S1972
- I1a2a1a1a1a2b4b1~ A376, S1957, S1973, S1982, Y6667, Y8338
- I1a2a1a1a1a2b4b1a~ A264/Y7521
- I1a2a1a1a1a2b4b1a1~ A681/Y7142
- I1a2a1a1a1a2b4b1a2~ A262
- I1a2a1a1a1a2b4b1b~ S2000
- I1a2a1a1a1a2b4b1b1~ S1961
- I1a2a1a1a1a2b4b1b2~ A6247
- I1a2a1a1a1a2b4b1c~ A6244
- I1a2a1a1a1a2b4b1d~ A6261
- I1a2a1a1a1a2b4b1e~ S2014
- I1a2a1a1a1a2b4b1a~ A264/Y7521
- I1a2a1a1a1a2b4b2~ A1156/BY400
- I1a2a1a1a1a2b4b1~ A376, S1957, S1973, S1982, Y6667, Y8338
- I1a2a1a1a1a2b5~ A5811
- I1a2a1a1a1a2c~ A12722, A1272 (all not included in Ancestry-Sample)
- I1a2a1a1a1a2a~ A1818/Y8335, A1830/Y14670, BY461/Y8333, BY462/Y8844
- I1a2a1a1a1a1 L338/S197
- I1a2a1a1a1b CTS10937/S4698/Z2538, CTS1148/S2190, CTS8996/S4699/Z2537, S2205, S10738, Y3146
- I1a2a1a1a1a S1954/YSC0000261, S1962^/Z4731^, S1968/Z4730, S1969, S1978^/Z4736^, S1987/Z4729, Z4727, Z4728
- I1a2a1a1a2 F2642.1/S2169.1 [F2642/S2169]
- I1a2a1a1a2~ AM00085/AMM442/CTS4963/DF76/S2163/V3243, AM00086/AMM443/CTS6739/S2166, AM00087/AMM444/CTS8691, AM00088/AMM445/CTS9191, AM00089/AMM446/CTS12971, AM00090/AMM447/S2155/Z2904
- I1a2a1a1a3 A196/Y5497
- I1a2a1a1a4~ A1374/BY477/Y15150
- I1a2a1a1a5~ FGC22406/Y6233, FGC22407/Y6232, Y6231
- I1a2a1a1a6~ A1603, A1604, A1605, A1606, A1607, A1610, A1616, A1618, A1620, A106991 (isogg.org/tree/ISOGG_HapgrpI.html).
"Haplogroup I-S1954 is descended from haplogroup I-M170. Among 23andMe research participants, haplogroup I-S1954 is commonly found among populations in the United Kingdom and Ireland. [...] Haplogroup I-S1954 is linked to Alexander Hamilton. [...I]n the 21st century, genealogists documented the paternal haplogroups of dozens of Hamilton's living descendants and concluded that the Founding Father's paternal haplogroup was a branch of I-DF29" (discover.23andme.com/haplogroup/I1a2a1a1a-paternal).
 1300 BCE - 700 CE: E/R related tree hypotheses for latest marker S1954/YSC0000261
1300 BCE - 700 CE: E/R related tree hypotheses for latest marker S1954/YSC0000261
"DNA Marker Index data for Marker: YSC0000261 on Chromosome: Y
(1) I-YSC0000261: YSC0000261.1 YSC0000261 S1954 YSC261.1 YSC261 rs751841691. Found in haplogroup I1a under Z2535 as an intermediate branch on ISOGG tree. See also YSC261.2 in haplogroup E1b. See also YSC261.3 in haplogroup R1b. See also YSC0000261.2 in haplogroup E1b. See also YSC0000261.3 in haplogroup R1b. See also YSC0000261.4 in haplogroup E1b. Phylogenetic Parent: Z2535. Phylogenetic Children: L338 (Thomas Krahn, FTDNA 2010), Pos. hg38: 12864559. Pos. hg19: 14976484.
(2) E-YSC0000261: YSC0000261.2 YSC261.2 rs751841691. Found in haplogroup E. See also YSC261.1 in haplogroup I1a. See also YSC261.3 in haplogroup R1b. See also YSC0000261.1 in haplogroup I1a. See also YSC0000261.3 in haplogroup R1b. See also YSC0000261.4 in haplogroup E1b. Coincident with Y161123 (YFull 2019). Pos. hg38: 12864559. Pos. hg19: 14976484" (genetichomeland.com/dna-marker/chromosome-Y/YSC0000261).
- E (M96)
- E1 (P147)
- E1b (P177)
- E-M5557
- E-M215
- E-M35
- E-L539
- E-M78
- E-Z22639
- E-V65
- E-PF2159
- E-Z21231
- E-Z21238
- E-CTS194
- E-Y30691
- E-V1174 HU162 * Y611144(T) * FT55933 +15 SNPs formed 2900 ybp, TMRCA 1300 ybp ids: Libya, Egypt, Marocco, Italy
- E-Y161123 Y161122 * Y161123 * YSC261/S1954/YSC0000261 formed 1300 ybp, TMRCA 150 ybp ids: Egypt (yfull.com/tree/E-V1174)
- E-V1174 HU162 * Y611144(T) * FT55933 +15 SNPs formed 2900 ybp, TMRCA 1300 ybp ids: Libya, Egypt, Marocco, Italy
- E-Y30691
- E-CTS194
- E-Z21238
- E-Z21231
- E-PF2159
- E-V65
- E-Z22639
- E-M78
- E-L539
- E-M35
- E-M215
- E-M5557
- E1b (P177)
- E1 (P147)
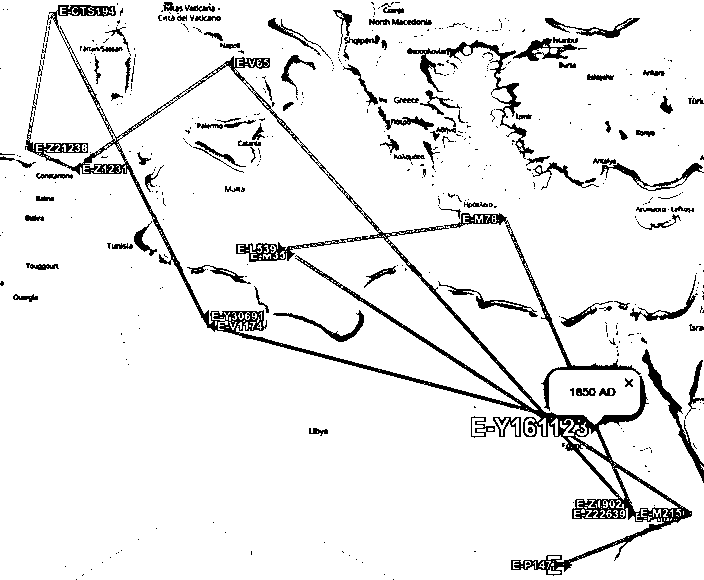
Abb. phylogeographer.com/snp-lookup/?I-YSC261, mygrations/?hg=E&clade=E-Y161123. Marker E-P147, E-P177, E-M215, E-M35 (8.000 BC), E-L539, E-M78, (5.000 BC, Zypern), E-V65 (1.500 BC, Süditalien), E-Z1231, E-Z21238, E-CTS194 (1.000 BC, Tunesien bis Sardinien), E-Y30691, E-V1174 (500 AD, Libyen), E-Y161123 (1.850 AD, Ägypten). Leaflet, Map data: OpenStreetMap contributors, CC-BY-SA, Imagery: Mapbox.
(3) "R-YSC0000261: YSC0000261.3 YSC261.3 rs751841691. Found in haplogroup R1b with S15337. See also YSC261.2 in haplogroup E1b. See also YSC261.1 in haplogroup I1a. See also YSC0000261.1 in haplogroup I1a. See also YSC0000261.2 in haplogroup E1b. See also YSC0000261.4 in haplogroup E1b. Coincident with BY76952 (YFull 2019). Pos. hg38: 12864559. Pos. hg19: 14976484" (genetichomeland.com/dna-marker/chromosome-Y/YSC0000261).
- F (M89)
- F1329 ("Defining mutation for haplogroup branch above HIJK. hg38 Ref does not match ancestral allele value. V2308 has alleles reversed")
- HIJK (M578)
- IJK (M523)
- K (M9)
- K2 (M526)
- K2b (M1221)
- P root on YFull and FTDNA trees (PF5850)
- P root (K2b2) on ISOGG tree (P295)
- M1254
- P337
- P1a (P284)
- P1a1 (P226)
- R (M207, "[o]riginated about 26,500 bce. Example is ancient sample MA1 from Irkutsk, Russia 22,000 bce", genetichomeland.com/welcome/dnapedigree.asp?RecordID=1708113)
- P1a1 (P226)
- P1a (P284)
- P337
- M1254
- P root (K2b2) on ISOGG tree (P295)
- P root on YFull and FTDNA trees (PF5850)
- K2b (M1221)
- K2 (M526)
- K (M9)
- IJK (M523)
- HIJK (M578)
- F1329 ("Defining mutation for haplogroup branch above HIJK. hg38 Ref does not match ancestral allele value. V2308 has alleles reversed")
- R (M207)
- Y482
- R1 (M173, "[s]o zeigten frühere Studien, dass in Europa vor allem die durch die Y-SNPs M173 und P25 charakterisierte Linie
R1b* vorliegt [Brión et al. 2005A]", WP)
- R1a ("[t]he highest frequency of R1a1a [M17, M198] in Europe is observed in Sorbs (63%), a West Slavic ethnic group, followed by Hungarians (60%)", "[a]ccording to Underhill et al. (2014), the downstream M417 (R1a1a1) subclade diversified into Z282 (R1a1a1b1a) and Z93 (R1a1a1b2) circa 5,800 years ago 'in the vicinity of Iran and Eastern Turkey'", "[a] genetic study published in Nature in March 2015 [Haak et al.] examined the remains of an Urnfield male buried in Halberstadt, Germany ca 1100-1000 BC. He was found to be a carrier of the paternal haplogroup R1a1a1b1a2 [S466/Z280, S204/Z91]", WP)
- R1b (M343, "Haplogruppe I1 sowie Untergruppen von R1b wie R1b-Z18 [R1b1a2a1a1b] und Untergruppen von R1a wie R1a-Z284 [R1a1a1b1a3, Isogg 2018] werden eng mit germanischen Völkern in Verbindung gebracht und sind mit den protogermanischen Sprechern der nordischen Bronzezeit verknüpft", WP zu I-M253, "Damgaard et al. (2018) analyzed the remains of a male and female buried at a Hallstatt cemetery near Litoměřice, Czech Republic between ca. 600 BC and 400 BC. The male carried paternal haplogroup R1b", WP)
- R1b1 (L754, P25)
- L761
- R1b1a (L389, M18, "R-L21 or R1b1a2a1a2c, also known as R-M529 or R-S145, is a Human Y-chromosome DNA haplogroup. It is often linked to the Insular Celts", WP. Other markers: S21, S28, M458, V13, DF27, Z2103, eupedia.com, 2018).
- R1b1a1 (P297)
- M269 ("Defining mutation for Western Atlantic Modal Haplotype (WAMH) of R1b in Europe. Originated about 11,000 bce")
- L23 ("Originated about 4,500 bce"; "Greco-Etruscan", eupedia.com/europe/genetische_Karten_von_Europa.shtml)
- L51 ("In Europe, almost entirely west of the Danube river")
- P310
- PF6538.1
- L151
- P312 ("Largest branch under haplogroup R1b. [...] Example is ancient sample esp005 from Spain 1,300 bce")
Continuation: Z46516, ZZ11_1, DF27, Z195, Z274, Z209, FGC83504, ZZ40_1, S21184.1, S19290.1, S16785.1, S15337, Y112817, BY11972, BY76952 ("Marker [YSC0000261.3] currently considered coincident with marker [BY76952], using that phylogenetic tree").
- P312 ("Largest branch under haplogroup R1b. [...] Example is ancient sample esp005 from Spain 1,300 bce")
- L151
- PF6538.1
- P310
- L51 ("In Europe, almost entirely west of the Danube river")
- L23 ("Originated about 4,500 bce"; "Greco-Etruscan", eupedia.com/europe/genetische_Karten_von_Europa.shtml)
- M269 ("Defining mutation for Western Atlantic Modal Haplotype (WAMH) of R1b in Europe. Originated about 11,000 bce")
- R1b1a1 (P297)
- R1b1b (M73)
- R1b1b2 ("In einer von Sims et al. [2007] publizierten Studie wurden drei bis dahin unbekannte, phylogenetisch informative Y-chromosomale SNP-Marker U152, U106 und U 198 mitgeteilt. Diese Marker repräsentieren die Haplogruppen (hg) R1b1b2h* (U152), R1b1b2g*, (U106) und R1b1b2g1 (U198)", M. Wirsching: "Vertiefte Analyse der Y-chromosomalen Linie R1b* in westdeutschen Populationen", Freiburg 2009, S. 15f.)
- R1b1a (L389, M18, "R-L21 or R1b1a2a1a2c, also known as R-M529 or R-S145, is a Human Y-chromosome DNA haplogroup. It is often linked to the Insular Celts", WP. Other markers: S21, S28, M458, V13, DF27, Z2103, eupedia.com, 2018).
- L761
- R1b1 (L754, P25)
- R1 (M173, "[s]o zeigten frühere Studien, dass in Europa vor allem die durch die Y-SNPs M173 und P25 charakterisierte Linie
R1b* vorliegt [Brión et al. 2005A]", WP)
- Y482
(4) "E-YSC0000261: YSC0000261.4 rs751841691. Found in haplogroup E1b. See also YSC0000261.1 in haplogroup I1a. See also YSC0000261.2 in haplogroup E1b. See also YSC0000261.3 in haplogroup R1b. Coincident with FTF58694 (FTDNA 2024). Pos. hg38: 12864559. Pos. hg19: 14976484" (genetichomeland.com/dna-marker/chromosome-Y/YSC0000261).
Zurück zum Inhaltsverzeichnis
E-Mail: kriswagenseil [at] gmx [point] de